pENTR4-H1 (#RDB04395)
Expression vector of RNAi driven by human H1 promoter.
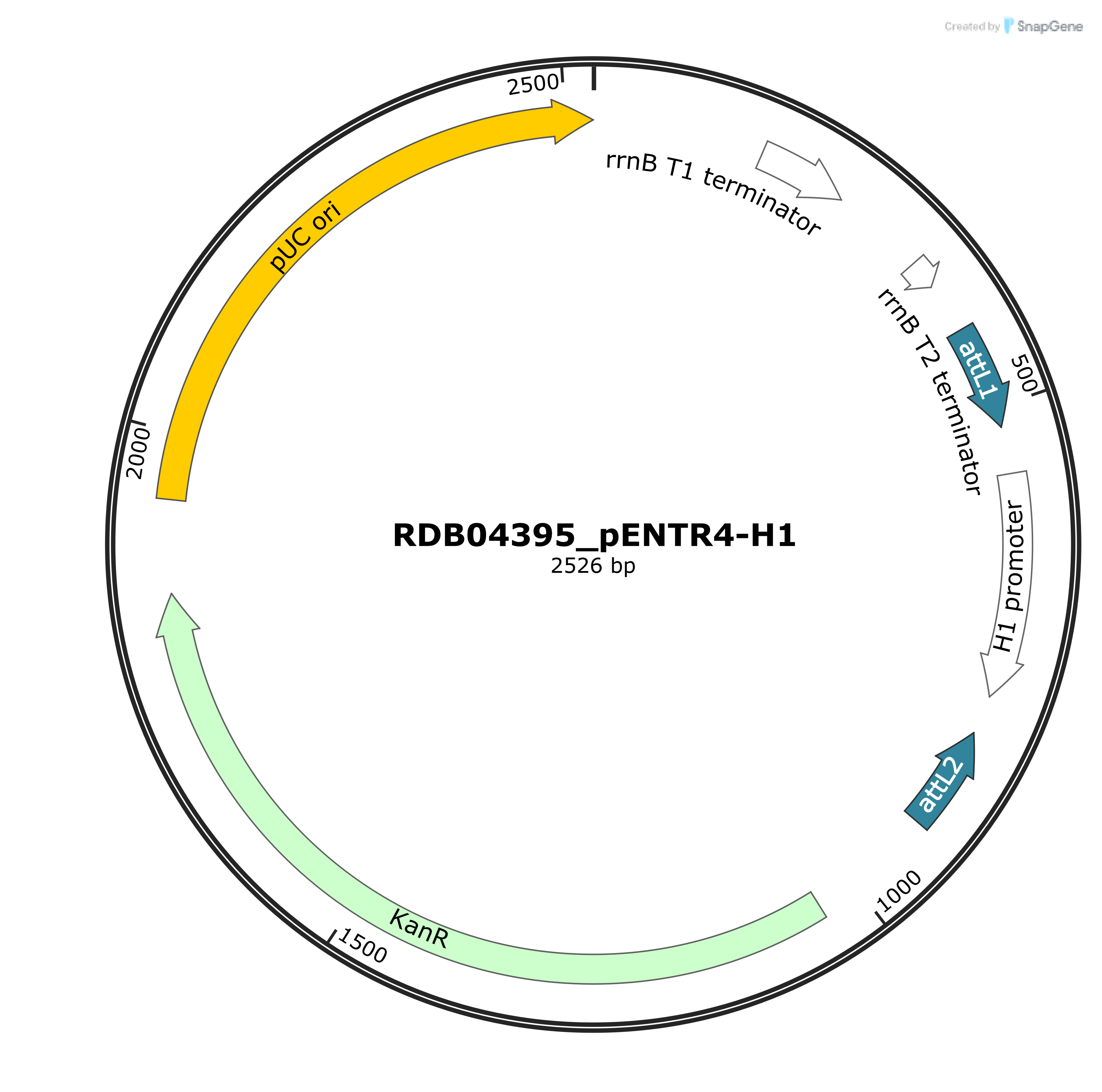 Drawn by SnapGene® software |
|
| Clone info. | Expression vector of RNAi driven by human H1 promoter. Frequently requested along with Gateway(R) destination vectors for constructing short hairpin RNA (shRNA) expressing vectors. |
|---|---|
| Vector backbone | pENTR4 (Plasmid) |
| Selectable markers | Kan^r |
| Growth conditions | 37C, LB+Kan |
| Gene/insert name | Human RPPH1 genomic DNA |
| Depositor|Developer | Miyoshi, Hiroyuki | |
Distribution information
Please check terms and conditions set forth by the depositor, which are specified in the RIKEN BRC Catalog and/or Web Catalog.
| Ordering forms | Order form [Credit Card Payment MTA, for use for not-for-profit academic purpose [Word Please visit Information of Request for Distribution.[link] |
|---|---|
| Terms and conditions for distribution | The availability of the BIOLOGICAL RESOURCE is limited to a RECIPIENT of a not-for profit institution for a not-for-profit academic purpose. The RECIPIENT agrees to expressly describe the late Dr. Hiroyuki Miyoshi as the Developer. |
 [open/close]
[open/close]| 必要書類 | 提供依頼書 提供同意書 (MTA, 非営利学術目的用)[Word 手続きの概要は、「提供申込みについて[link]」をご覧ください。 |
|---|---|
| MTAに書く使用条件 | 本件研究材料は、非営利機関の非営利学術研究に限って提供する。本件研究材料を利用した研究結果等を発表する際は、本件研究材料が故三好浩之博士により開発されたことを明示する。 |
| Catalog # | Resource name | Shipping form | Fee |
|---|---|---|---|
| RDB04395 | pENTR4-H1 | DNA solution |
JPY 9,460 (not-for-profit academic purpose) plus cost of shipping containers, dry ice (if required) and shipping charge |
How to cite this biological resource
Materials & Methods section:
| The pENTR4-H1 was provided by the RIKEN BRC through the National BioResource Project of the MEXT, Japan (cat. RDB04395). |
Reference section:
Further references such as user reports and related articles (go to bottom)
References
Original, user report and related articles
| user_report | Yamagishi, M., Mechanisms of action and resistance in histone methylation-targeted therapy. Nature 627 (8002): 221-228 (2024). PMID 38383791. [link to RRC of NBRP] |
|---|---|
| user_report | Matsumoto, R., AP-1B regulates interactions of epithelial cells and intraepithelial lymphocytes in the intestine. Cell. Mol. Life Sci. 81 (1): 425 (2024). PMID 39369131. [link to RRC of NBRP] |
| user_report | Yang, S., Neutrophil Extracellular Traps Delay Diabetic Wound Healing by Inducing Endothelial-to-Mesenchymal Transition via the Hippo pathway. Int J Biol Sci. 2023 Jan 1;19(1):347-361 (2023). PMID 36594092. [link to RRC of NBRP] |
| user_report | Araki, H., LSD1 defines the fiber type-selective responsiveness to environmental stress in skeletal muscle. Elife 12: e84618 (2023). PMID 36695573. [link to RRC of NBRP] |
| user_report | Watanabe, H., Astrocytic APOE4 genotype-mediated negative impacts on synaptic architecture in human pluripotent stem cell model. Stem Cell Reports 18 (9): 1854-1869 (2023). PMID 37657448. [link to RRC of NBRP] |
| user_report | Truong, T.T.K., Arl4c is involved in tooth germ development through osteoblastic/ameloblastic differentiation. Biochem. Biophys. Res. Commun. 679: 167-174 (2023). PMID 37703759. [link to RRC of NBRP] |
| user_report | Shimizu, K., Diurnal variation in declarative memory and the involvement of SCOP in cognitive functions in nonhuman primates. 02 November 2022, PREPRINT (Version 1) available at Research Square https://doi.org/10.21203/rs.3.rs-2013762/v1 [link to Journal] |
| user_report | Miyakuni, K., Genome-wide analysis of DNA methylation identifies the apoptosis-related gene UQCRH as a tumor suppressor in renal cancer. Mol. Oncol. 16 (3): 732-749 (2022). PMID 34133843. [link to RRC of NBRP] |
| user_report | Yamashita, A., Culture substrate-associated YAP inactivation underlies chondrogenic differentiation of human induced pluripotent stem cells. Stem Cells Transl. Med. 10 (1): 115-127 (2021). PMID 32822104. [link to RRC of NBRP] |
| user_report | Funaki, T., CADM1 promotes malignant features of small-cell lung cancer by recruiting 4.1R to the plasma membrane. Biochem. Biophys. Res. Commun. 534: 172-178 (2021). PMID 33298314. [link to RRC of NBRP] |
| user_report | Totani, H., Autocrine HGF/c-Met signaling pathway confers aggressiveness in lymph node adult T-cell leukemia/lymphoma. Oncogene 39 (35): 5782-5794 (2020). PMID 32747750. [link to RRC of NBRP] |
| user_report | Yuki, R., Desuppression of TGF-beta signaling via nuclear c-Abl-mediated phosphorylation of TIF1 gamma/TRIM33 at Tyr-524, -610, and -1048. Oncogene 38 (5): 637-655 (2019). PMID 30177833. [link to RRC of NBRP] |
| user_report | Ito, T., Mathematical analysis of gefitinib resistance of lung adenocarcinoma caused by MET amplification. Biochem. Biophys. Res. Commun. 511 (3): 544-550 (2019). PMID 30824185. [link to RRC of NBRP] |
| user_report | Nakajo, H., Role for tyrosine phosphorylation of SUV39H1 histone methyltransferase in enhanced trimethylation of histone H3K9 via neuregulin-1/ErbB4 nuclear signaling. Biochem. Biophys. Res. Commun. 511 (4): 765-771 (2019). PMID 30833073. [link to RRC of NBRP] |
| user_report | Kamil, M., High filamin-C expression predicts enhanced invasiveness and poor outcome in glioblastoma multiforme. Br. J. Cancer. 120 (8): 819-826 (2019). PMID 30867563. [link to RRC of NBRP] |
| user_report | Makino, Y., Single cell RNA-sequencing identified Dec2 as a suppressive factor for spermatogonial differentiation by inhibiting Sohlh1 expression. Sci. Rep. 9 (1): 6063 (2019). PMID 30988352. [link to RRC of NBRP] |
| user_report | Yamagishi, M., Targeting Excessive EZH1 and EZH2 Activities for Abnormal Histone Methylation and Transcription Network in Malignant Lymphomas. Cell Rep. 29 (8): 2321-2337.e7 (2019). PMID 31747604. [link to RRC of NBRP] |
| user_report | Jinnou, H., Radial Glial Fibers Promote Neuronal Migration and Functional Recovery after Neonatal Brain Injury. Cell Stem Cell 22 (1): 128-137.e9. (2018). PMID 29276142. [link to RRC of NBRP] |
| user_report | Kawasaki, N., TUFT1 interacts with RABGAP1 and regulates mTORC1 signaling Cell Discovery 4 (1): (2018). PMID 29423269. [link to RRC of NBRP] |
| user_report | Funato, K., SIRT2-mediated inactivation of p73 is required for glioblastoma tumorigenicity. EMBO Rep. 19 (11). pii: e45587 (2018). PMID 30213795. [link to RRC of NBRP] |
| user_report | Neri, S., Fibroblast-led cancer cell invasion is activated by epithelial-mesenchymal transition through platelet-derived growth factor BB secretion of lung adenocarcinoma. Cancer Lett. 395: 20-30 (2017). PMID 28286261. [link to RRC of NBRP] |
| user_report | Ishibashi, M., CD200-positive cancer associated fibroblasts augment the sensitivity of Epidermal Growth Factor Receptor mutation-positive lung adenocarcinomas to EGFR Tyrosine kinase inhibitors. Sci. Rep. 7: 46662 (2017). PMID 28429795. [link to RRC of NBRP] |
| user_report | Katsura, A., MicroRNA-31 is a positive modulator of endothelial-mesenchymal transition and associated secretory phenotype induced by TGF-beta. Genes Cells 21 (1): 99-116 PMID 26663584. [link to RRC of NBRP] |
| user_report | Takahashi, Y., Perilipin2 plays a positive role in adipocytes during lipolysis by escaping proteasomal degradation. Sci Rep. 6: 20975 (2016). PMID 26876687. [link to RRC of NBRP] |
| user_report | Hikasa, H., Merlin/NF2-Lin28B-let-7 Is a Tumor-Suppressive Pathway that Is Cell-Density Dependent and Hippo Independent. Cell Rep. 14 (12): 2950-2961 (2016). PMID 26997273. [link to RRC of NBRP] |
| user_report | Johmura, Y., Defective DNA repair increases susceptibility to senescence through extension of Chk1-mediated G2 checkpoint activation. Sci. Rep. 6: 31194 (2016). PMID 27507734. [link to RRC of NBRP] |
| user_report | Inoue, J., Identification of BCL11B as a regulator of adipogenesis. Sci. Rep. 6: 32750 (2016). PMID 27586877. [link to RRC of NBRP] |
| user_report | Matsuda, Y., Epigenetic Heterogeneity in HIV-1 Latency Establishment. Sci. Rep. 5: 7701 (2015). PMID 25572573. [link to RRC of NBRP] |
| user_report | Neri, S., Podoplanin-expressing cancer-associated fibroblasts lead and enhance the local invasion of cancer cells in lung adenocarcinoma. Int. J. Cancer. 137 (4): 784-796 (2015). PMID 25648219. [link to RRC of NBRP] |
| user_report | Yamagishi M, Polycomb-Mediated Loss of miR-31 Activates NIK-Dependent NF-κB Pathway in Adult T Cell Leukemia and Other Cancers. Cancer Cell, 21 (1): 121-135 (2012). PMID 22264793. [link to RRC of NBRP] |
| user_report | Sasaki, M., Equine major histocompatibility complex class I molecules act as entry receptors that bind to equine herpesvirus-1 glycoprotein D. Genes Cells, 16 (4), 343-357 (2011). PMID 21306483. [link to RRC of NBRP] |
| user_report | Aoyama, K., Nuclear c-Abl-mediated tyrosine phosphorylation induces chromatin structural changes through histone modifications that include H4K16 hypoacetylation. Exp. Cell Res., 317 (20): 2874-903 (2011). PMID 22001646. [link to RRC of NBRP] |
| user_report | Tanaka, Y., Dual Function of Histone H3 Lysine 36 Methyltransferase ASH1 in Regulation of Hox Gene Expression. PLoS One., 6 (11): e28171 (2011). PMID 22140534. [link to RRC of NBRP] |
| user_report | Ohoka, N., The orphan nuclear receptor RORalpha restrains adipocyte differentiation through a reduction of C/EBPbeta activity and perilipin gene expression. Mol. Endocrinol., 23, 759-771 (2009). PMID 19324970. [link to RRC of NBRP] |
| user_report | Nakayama Y., Bleomycin-induced over-replication involves sustained inhibition of mitotic entry through the ATM/ATR pathway. Exp. Cell Res., 315, 2515-2528 (2009). PMID 19527713. [link to RRC of NBRP] |
| user_report | Tanabe, R., PRRX1 induced by BMP signaling decreases tumorigenesis by epigenetically regulating glioma-initiating cell properties via DNA methyltransferase 3A. Mol. Oncol. 16 (1): 269-288 (2022). PMID 34214250. [link to RRC of NBRP] |
Resource search
Link
Other resources in our bankExternal Database
Human RPPH1
Reference sequence
Tips
Featured content
- Empty Backbone (English text)
- Empty Backbone (Japanese text)
- Lentivirus Vector (English text)
- Lentivirus Vector (Japanese text)




